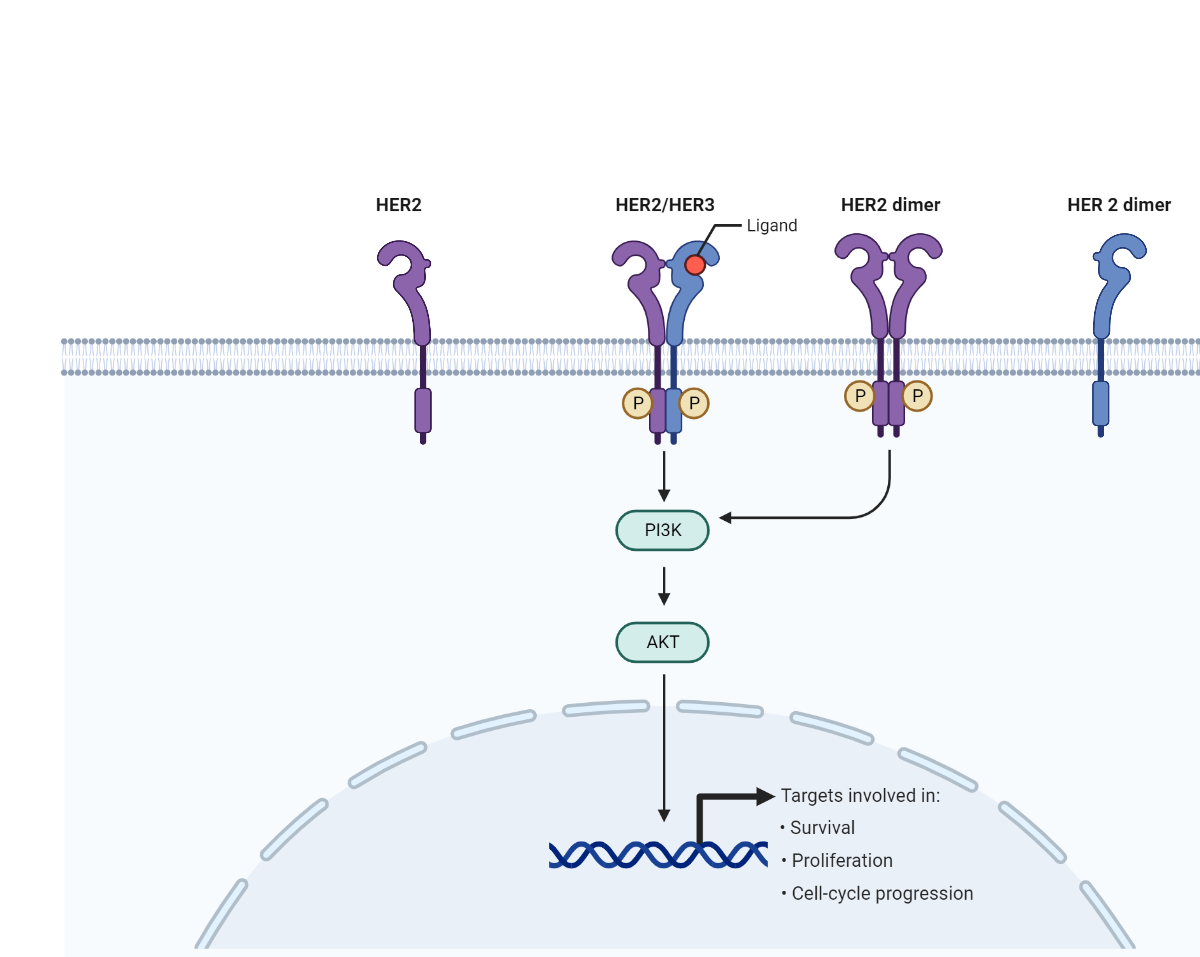
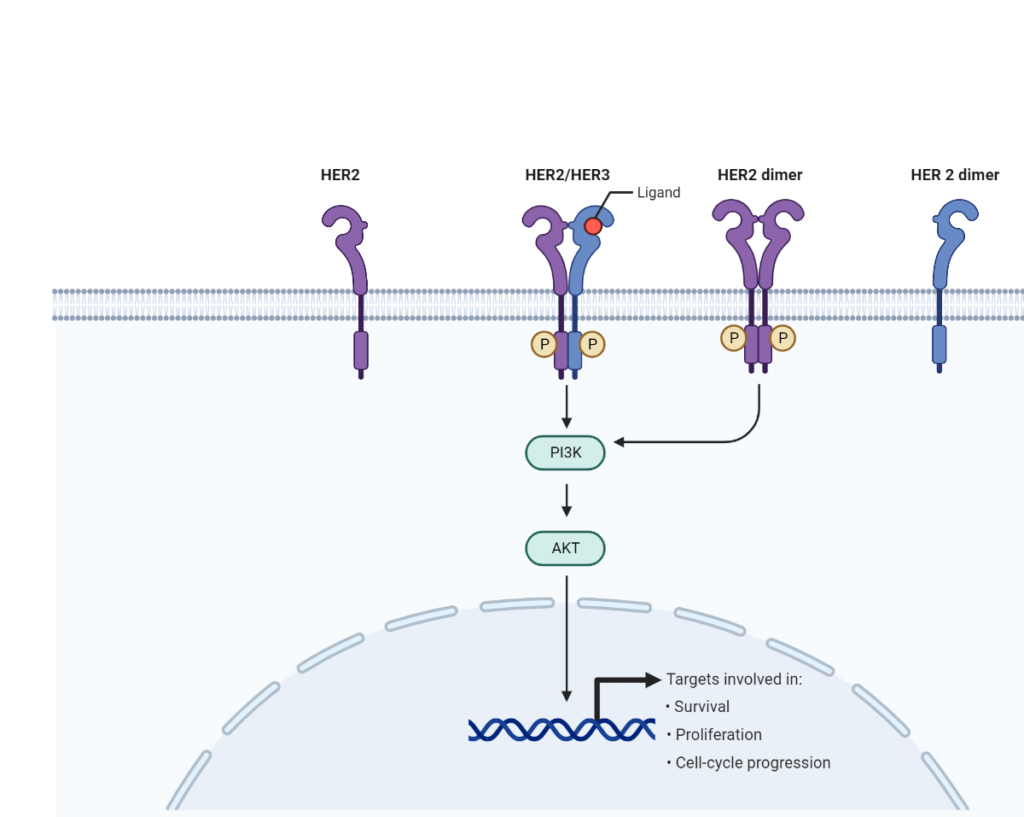
Detecting activated biomarker signals is challenging with traditional IHC
In clinical oncology, immunohistochemistry (IHC) is often used to determine the presence or absence of protein biomarkers. Clinical scientists will section and mount tumor samples, then stain using a primary antibody for the target biomarker. Because detection is done on a single section of tissue at a time, biomarkers can be spatially localized to regions of tumor within the tissue. However, standard immunohistochemical processes, such a formalin fixation, make some protein biomarker epitopes difficult to detect, particularly phosphorylation. As phosphorylation is often a signal for protein activation, this means oncologists are missing out on a powerful source of information regarding tumor activity. The solution? Reverse Phase Protein Arrays (RPPA).
RPPA detects phosphorylation with ease
In RPPA, cellular proteins are instead enriched from a tumor epithelium via laser capture microdissection and denatured in lysis buffer. These enriched tumor samples are then spotted onto slides, probed by a primary antibody, and visualized by a fluorescent marker. From only a few sections of a core needle biopsy, upwards of 100 cellular proteins can be analyzed. Unlike IHC, the RPPA methodology allows for analysis of phosphorylated proteins, including samples that were fixed in formalin or fresh-frozen. This grants oncologists deeper insights into selecting an appropriate therapy that would otherwise be lost with IHC.
Let’s take the HER2 pathway as an example in the importance of phosphorylation
HER2 is a receptor tyrosine-protein kinase commonly associated with breast cancer. HER2 resides in the cell membrane, where it can dimerize with other HER proteins (like HER3). The dimerized proteins then become phosphorylated and stimulate cell growth and proliferation. Oncologists currently use IHC to classify breast tumors for HER2 activity using a 0-3 point system, with 0 to 1+ representing HER2 negative tumors, 2+ representing a borderline case, and 3+ representing HER2 positive tumors. If a tumor is HER2+, oncologists may decide to prescribe patients with HER2 inhibitors as a mode of therapy.
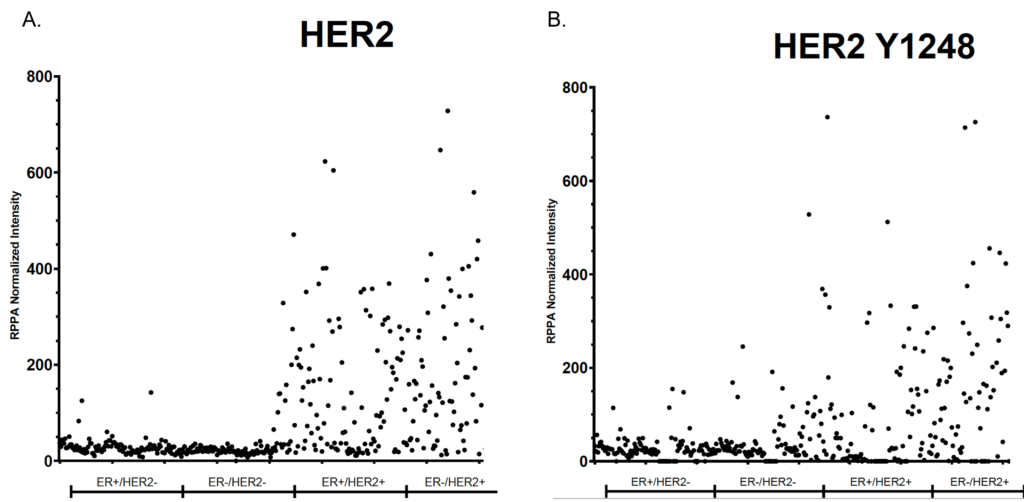
RPPA reveals some patients classified as HER2- have high phosphorylated HER2 levels
When we compare IHC to RPPA in measuring total HER2 in tumor tissue, in general we see high agreement, with samples classified “HER2+” with IHC having high HER2 levels in the RPPA assay (Figure 1a). However, if we look at HER2 Y1248 phosphorylation using RPPA, we can see that some samples called “HER2-” actually have high levels of the phosphorylated HER2 protein (Figure 1b).
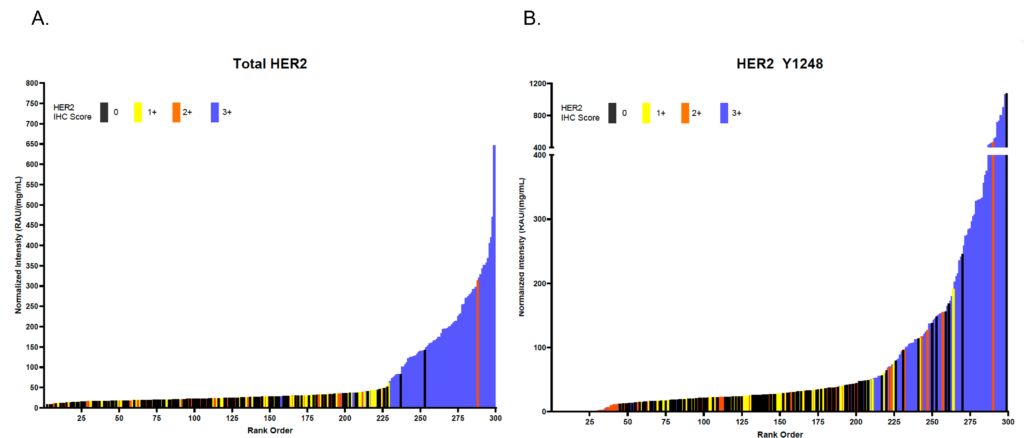
Looking at the data from a different angle, we can rank samples based on their RPPA signal strength, and color code based on IHC score. Again, we see when looking at HER2 total levels there is a strong concordance between the methods (Figure 2A). However, when we look at HER2 phosphorylation levels, we see samples with high HER2 phosphorylation start becoming mixed in their IHC score, suggesting IHC alone may not be sufficient in determining who will respond well to HER2 inhibitors (Figure 2B). This means some patients may benefit from HER2 inhibitors even though they are clinically classified HER2-.
Real world results emphasize the need for higher-resolution tests
Based on all these advantages, researchers are now turning to RPPA for better patient insights. Clark et al. (2021) found that by measuring phosphorylation levels of HER2 and HER3, researchers could predict pathologic Complete Response (pCR) independent of HER2 IHC data. Importantly, this included patients who were HER2+ based on IHC but did not achieve pCR, as well as the converse case (HER2- but did achieve pCR). Moving forward, Theralink will continue to support these types of research that furthers precision medicine. We currently can probe for over 700 protein targets (browse them and their pathways here), including many phosphorylated states, with more being added all the time. Contact one of our scientists today to learn how we can support your work.